Most biological systems consist of many moving parts that happen at slightly different timescales. A true-to-life biological model of a sub-network might include dozens of parameters and biological species, regardless of the scale. Constructing a model with an appropriate level of complexity is one of the challenges in mathematical biology.
Why is model complexity an issue?
Pros: A more complex model is easier to interpret, incorporates interesting biological dynamics, minimizes the amount of biological assumptions that are made by the modeler.
Cons: Complex models include more parameters, making the dynamics more heavily reliant on optimization. Model fit is not an easy thing to determine for complex models, and the benefit (mathematically speaking) may not be quantifiable.
Background
From statistical modeling we know that there is a bias-variance trade-off:
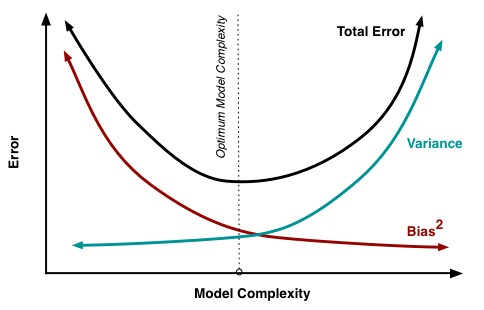
Variance is the degree to which our model fit is specific to the current data set. If your model was applied to another data set, how difficult would it be to get it to fit? Bias can be thought of as the degree to which our model fits the current data set. Theoretically, if you have a poor fit to your current data, you will have a roughly equally poor fit to new data. Model complexity affects both of these values.
Model complexity can be considered as coming from two sources:
- Parameters
- Variables or components
Parameters
In statistical modeling, model complexity is typically understood as the number of parameters included in the model. Complex models may also include interactions (nonlinear effects) between the variables or components within the model. Biological systems with multiple components with unique dynamics have an additional layer of complexity.
Components
Biological systems normally consist of many components. However, these components are usually reducible or semi-redundant, leaving room for simplification. It isn’t clear how additional components directly affects model complexity (as far as I know), but generally additional components require additional parameters.
Model simplification techniques
For a biological system of equations with multiple components and parameters, there are 3 ways to simplify the model:
- Parameter Reduction – Reducing the number of parameters is the most clear option for simplifying a model. This results in fewer values to optimize during curve-fitting, as well as less model flexibility. Besides removing biological effects from your model, you might also consider two parameters to be the same effective rate, and reduce the total number in that way.
- Component Reduction – Reducing the number of components is typically not preferred for mathematical biology. To justify this reduction, you can perform sensitivity analysis to determine components that don’t affect the dynamics significantly.
- Equation Simplification – The manner in which the equations in the model are constructed greatly effects simulation dynamics. Equation writing strategies may also be helpful in reducing parameter number. However, it isn’t clear how “simple” equations affect the complexity of the system of equations, particularly if there is no change in the number of parameters or components. Keep in mind these strategies reduce the interpretability of the model.
